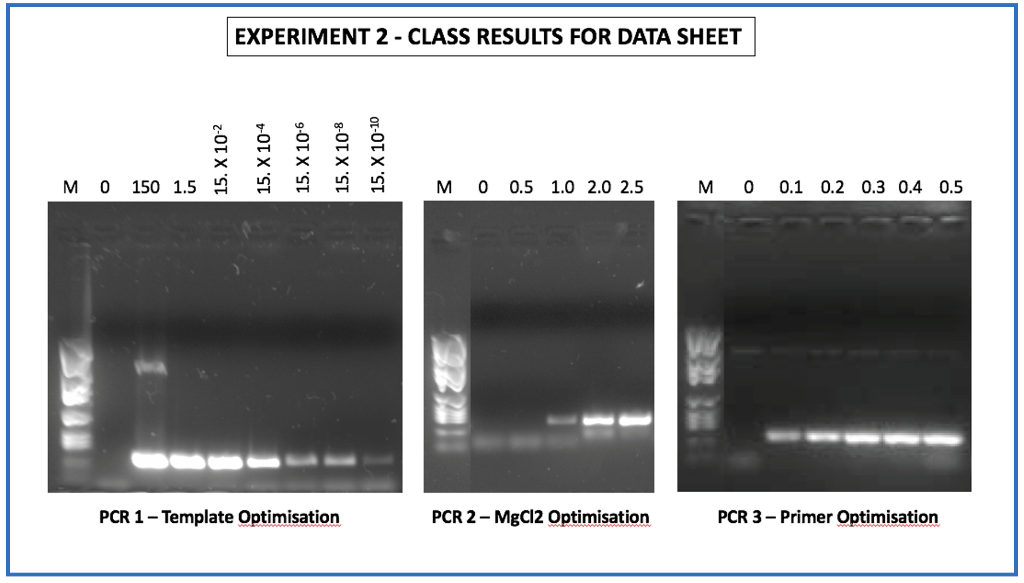
An optimized protocol for stepwise optimization of real-time RT-PCR analysis | Horticulture Research

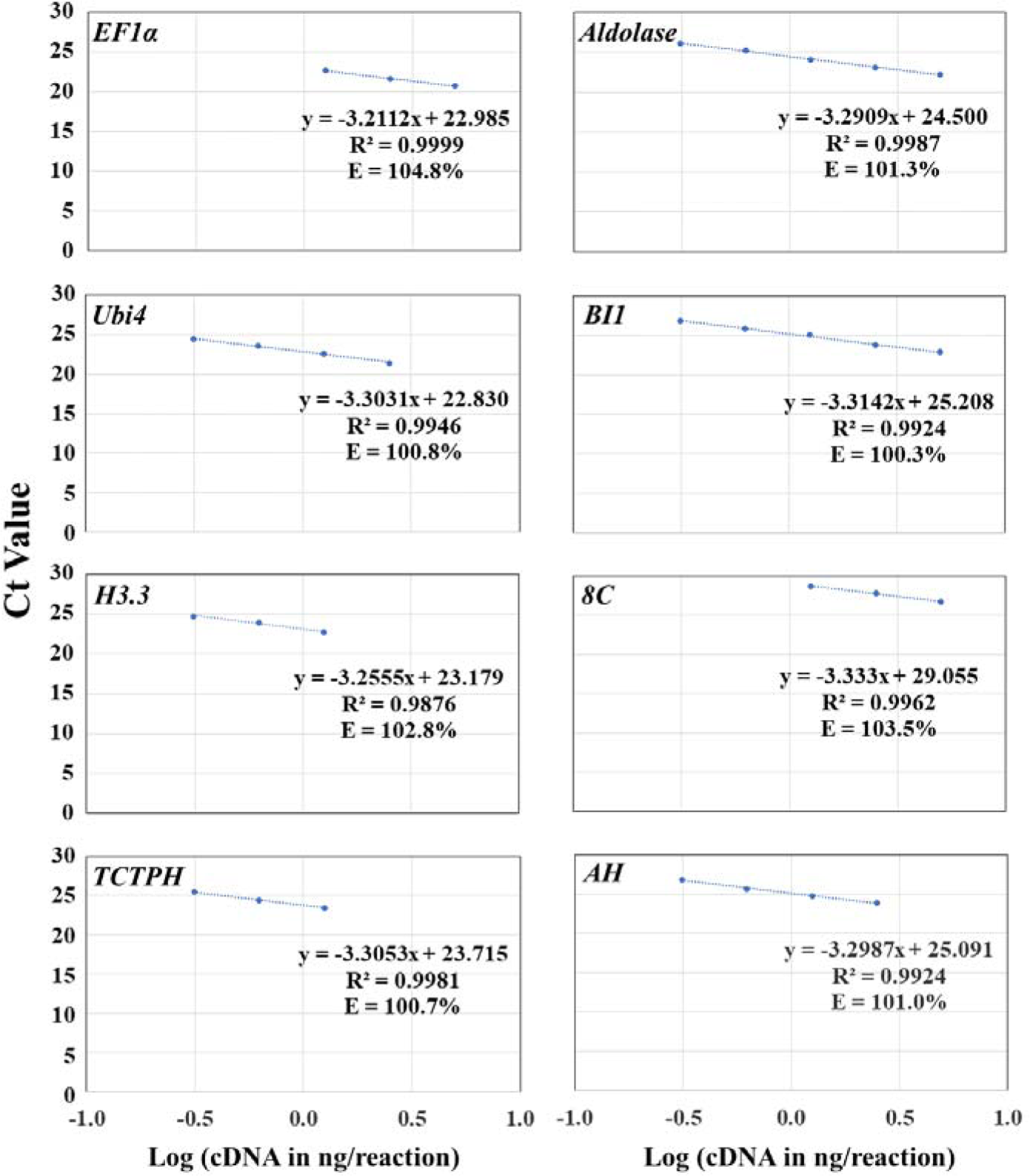
Optimizations of primers and EvaGreen concentrations. To provide an... | Download Scientific Diagram
A novel metric to improve mismatched primer selection and quantification accuracy in amplifying DNA repeats for quantitative polymerase chain reactions | PLOS ONE

Figure 2 from COMplementary Primer ASymmetric PCR (COMPAS-PCR) Applied to the Identification of Salmo salar, Salmo trutta and Their Hybrids | Semantic Scholar

SOLVED: I need to calculate the primer concentration after PCR cycle. My DNA segment (798 bp) is amplified by PCR. The 50 uL mixture of PCR reaction. Primer concentration is 2 uM.

Diagnostics | Free Full-Text | Thermostable Heptaplex PCR Assay for the Detection of Six Respiratory Bacterial Pathogens

SOLVED: 1. 2.5 μl of 10 μM working primer concentration is added to a 47.5 μl PCR reaction. What is the final concentration of primer in the PCR reaction? 2. Is this
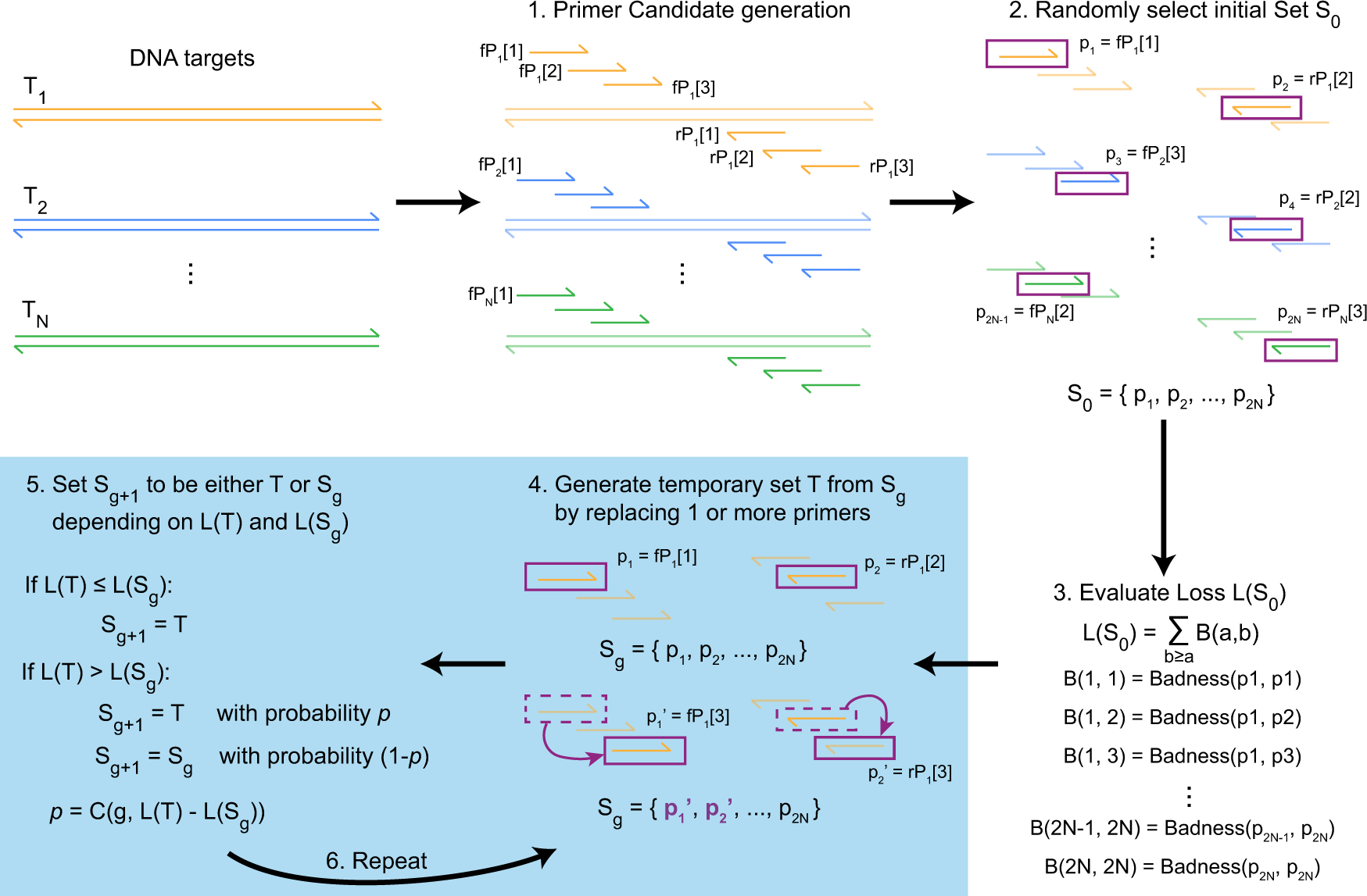
Designing highly multiplex PCR primer sets with Simulated Annealing Design using Dimer Likelihood Estimation (SADDLE) | Nature Communications

Optimization of universal primer set for the identification of all Influenza A viruses using real-time PCR | bioRxiv

Unreacted Labeled PCR Primers Inhibit the Signal in a Nucleic Acid Lateral Flow Assay as Elucidated by a Transport Reaction Model | ACS Measurement Science Au

Highly multiplex PCR assays by coupling the 5′-flap endonuclease activity of Taq DNA polymerase and molecular beacon reporters | PNAS